Smoker Status Prediction Using Edge List Table
This example is based on the Smoker Status Prediction dataset from Kaggle.
In this example, we use the RelationalAI Predictive Reasoner to predict whether a person is a smoker or not—a binary classification task.
The data is represented using a database of two tables: PEOPLE and RELATED.
PEOPLE: Each row in thePEOPLEtable represents an individual, along with their demographic and medical attributes such as age, height, cholesterol, triglyceride levels, and more. TheIdcolumn uniquely identifies each person and serves as the candidate key. Below are the first five records from the table, illustrating its structure and sample values.
| Id | age | height (cm) | weight (kg) | systolic | relaxation | fasting blood sugar | Cholesterol | triglyceride | HDL | LDL | hemoglobin | Urine protein | serum creatinine | AST | ALT | Gtp |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 35 | 170 | 85 | 118 | 78 | 97 | 239 | 153 | 70 | 142 | 19.8 | 1 | 1.0 | 61 | 115 | 125 |
| 2 | 20 | 175 | 110 | 119 | 79 | 88 | 211 | 128 | 71 | 114 | 15.9 | 1 | 1.1 | 19 | 25 | 30 |
| 3 | 45 | 155 | 65 | 110 | 80 | 80 | 193 | 120 | 57 | 112 | 13.7 | 3 | 0.6 | 1090 | 1400 | 276 |
| 4 | 45 | 165 | 80 | 158 | 88 | 249 | 210 | 366 | 46 | 91 | 16.9 | 1 | 0.9 | 32 | 36 | 36 |
| 5 | 20 | 165 | 60 | 109 | 64 | 100 | 179 | 200 | 47 | 92 | 14.9 | 1 | 1.2 | 26 | 28 | 15 |
RELATED: TheRELATEDtable encodes relationships between individuals. Each row contains two columns—person1andperson2—both referencing theIdcolumn in thePEOPLEtable. These are used as foreign keys. This table was generated by randomly pairing individuals, with a higher probability assigned to pairs where both individuals share the same smoking status (either both smokers or both non-smokers). Since this table has two columns and both are foreign keys, this table can be represented with an edge list. Below are the first five records from the table.
| Person1 | Person2 |
|---|---|
| 1 | 24042 |
| 1 | 25899 |
| 1 | 37454 |
| 2 | 10200 |
| 2 | 13384 |
The goal is to leverage the relational structure of this data to predict whether each person is a smoker, using both individual features and the network of relationships captured in the edge list.
For that purpose we create train, validation, and test tables, each including:
- An
Idcolumn that identifies the person whose smoking status the model is trying to predict. - A
smokingcolumn containing the binary label for prediction:1indicates that the person smokes, and0indicates they do not. This column is not required in the test table, as it is only used for model training and evaluation (e.g., calculating metrics like accuracy or F1 score), while the test table itself is solely intended for generating predictions.
Below are the first five rows of the train table:
| Id | Smoking |
|---|---|
| 23109 | 1 |
| 6421 | 0 |
| 24852 | 0 |
| 13773 | 0 |
| 15060 | 0 |
You can download all the tables required to run this experiment from the links below:
Create a Provider and a SnowflakeConnector
Section titled “Create a Provider and a SnowflakeConnector”Create a Provider.
from relationalai_gnns import Provider
snowflake_config = { "user": "<SNOWFLAKE_USER>", "password": "<SNOWFLAKE_PASSWORD>", "account": "<SNOWFLAKE_ACCOUNT>", "role":"<SNOWFLAKE_ROLE_WITH_ACCESS_TO_DB>", "warehouse": "<SNOWFLAKE_WAREHOUSE>", "app_name": "<RELATIONALAI_APP_NAME>", "auth_method": "password"}provider = Provider(**snowflake_config)from relationalai_gnns import Provider
snowflake_config = { "app_name": "<RELATIONALAI_APP_NAME>", "auth_method": "active_session", "role": "<SNOWFLAKE_ROLE_WITH_ACCESS_TO_DB>"}provider = Provider(**snowflake_config)Create a GNN reasoner.
provider.create_gnn( name="my_gnn_reasoner", size="HIGHMEM_X64_S")Create a SnowflakeConnector.
from relationalai_gnns import SnowflakeConnector
connector = SnowflakeConnector( **snowflake_config, engine_name = "my_gnn_reasoner")Define Nodes and Edges from the Snowfake Tables
Section titled “Define Nodes and Edges from the Snowfake Tables”The rows of the PEOPLE table are represented as nodes by defining a GNNTable of type node with the candidate key Id.
The rows of the RELATED table are represented as edges connecting the people nodes, defined as a GNNTable of type edge with two foreign keys—Person1 and Person2—both referencing the Id column of the PEOPLE table.
Since the RELATED table contains only these two columns, each corresponding to a foreign key, it naturally represents an edge in the graph.
Essentially, the combination of the node GNNTable (PEOPLE) and the edge GNNTable (RELATED) enables us to model a graph of relationships between different people, where each person is a node and each relationship forms an edge between them.
from relationalai_gnns import GNNTable, ForeignKey, CandidateKey
# Create a node GNNTable from the 'PEOPLE' table from 'SMOKERS_DB.DATA' schema in Snowflake,# with candidate key column named 'Id'.people_table = GNNTable( connector=connector, name="People", type="node", source="SMOKERS_DB.DATA.PEOPLE", candidate_keys=[CandidateKey(column_name="Id")],)
# Create an edge from the 'RELATED' table from 'SMOKERS_DB.DATA' schema in Snowflake.# Add two foreign keys:# Both referencing People.Id,# where People is the name of the "SMOKERS_DB.DATA.PEOPLE" table defined above,# and Id is its candidate key column.related_table = GNNTable( connector=connector, name="Related", source="SMOKERS_DB.DATA.RELATED", type="edge", foreign_keys=[ ForeignKey(column_name="person1", link_to="People.Id"), ForeignKey(column_name="person2", link_to="People.Id"), ],)Define the learning task
Section titled “Define the learning task”We then define a NodeTask for binary classification. You will need to prepare three tables with class labels:
SMOKERS_DB.TASK.TRAINSMOKERS_DB.TASK.VALIDATIONSMOKERS_DB.TASK.TEST
Each table must include:
- An
Idcolumn that serves as a foreign key referencing thePeopleGNNTable. - A
smokingcolumn containing the binary label for prediction:1indicates that the person smokes, and0indicates that they do not. This column is not necessary for the test table.
from relationalai_gnns import NodeTask, TaskType, ForeignKey
node_task = NodeTask( connector=connector, name="smoking", task_data_source={ "train": SMOKERS_DB.TASK.TRAIN, "test": SMOKERS_DB.TASK.TEST, "validation": SMOKERS_DB.TASK.VALIDATION, }, target_entity_column=ForeignKey(column_name="Id", link_to="People.Id"), label_column="smoking", task_type=TaskType.BINARY_CLASSIFICATION,)node_task.set_evaluation_metric(EvaluationMetric(name="roc_auc"))In the definition of the node_task:
- The
target_entity_columnspecifies the IDs of the individuals for whom predictions are to be made. - The evaluation metric used to determine the best model is set to ROC AUC.
Compose a Dataset from GNNTables and a Task
Section titled “Compose a Dataset from GNNTables and a Task”Construct a Dataset as follows:
from relationalai_gnns import Dataset
dataset = Dataset( connector=connector, dataset_name="smoking", tables=[ people_table, related_table, ], task_description=node_task,)The .visualize_dataset() method is provided to help you inspect the connectivity between all your tables and the task.
from IPython.display import Image, display
graph = dataset.visualize_dataset()plt = Image(graph.create_png())display(plt)from graphviz import Source
graph = dataset.visualize_dataset()# Experiment with font size and plot size to get a good visualizationfor node in graph.get_nodes(): font_size = node.get_attributes()['fontsize'] font_size = "16" node.set('fontsize', font_size)
graph.set_graph_defaults(size="10,10!") # Increase graph size
src = Source(graph.to_string())src # Display in notebook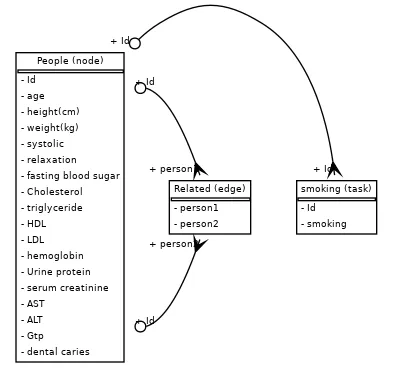
Train a model and generate predictions
Section titled “Train a model and generate predictions”Once your Dataset is ready, you can proceed to train a model.
from relationalai_gnns import TrainerConfig, Trainer
trainer_config = TrainerConfig(connector=connector, device="cuda", n_epochs=15)
trainer = Trainer(connector=connector, config=trainer_config)
train_job = trainer.fit(dataset=dataset)You can use get_status() to check the progress and performance of the training job:
train_job.get_status()After training has finished it returns information such as:
{'job_id': '88fad2a7-e128-4171-88c8-180518363fba', 'status': 'COMPLETED', 'started_at': '2025-10-16T15:41:07.459188', 'finished_at': '2025-10-16T15:55:13.667735', 'updated_at': '2025-10-16T15:55:13.667739', 'experiment_name': 'smoking_binary_classification_smoking', 'job_type': 'train', 'model_run_id': 'e506d38321f2411fb86179a3cef94988', 'result': {'metrics': {'1': {'train_loss': 0.16520570157509146, 'val_loss': 0.032000904575371594, 'val_metrics': {'average_precision': 0.9992711360570745, 'accuracy': 0.9892252437147255, 'f1': 0.985252808988764, 'roc_auc': 0.9995749455160009}, 'train_metrics': {'average_precision': 0.9721073764220794, 'accuracy': 0.9246480905505499, 'f1': 0.8925468678555099, 'roc_auc': 0.9816292750050081}, 'tune_metric': 'roc_auc'}, ... '5': {'train_loss': 0.007102684612607561, 'val_loss': 0.006133774527536999, 'val_metrics': {'average_precision': 0.9999741820696594, 'accuracy': 0.9976911236531555, 'f1': 0.9968542467668647, 'roc_auc': 0.999984991413956}, 'train_metrics': {'average_precision': 0.9999490578983604, 'accuracy': 0.9976913457530381, 'f1': 0.9968561697668326, 'roc_auc': 0.9999706709683405}, 'tune_metric': 'roc_auc'}, ... }'best_epoch': 5}}Once the model is trained, you can generate batch predictions using the predict() method:
from relationalai_gnns import OutputConfig
inference_job = trainer.predict( output_alias="experiment_1", output_config=OutputConfig.snowflake( database_name="SMOKERS_DB", schema_name="PUBLIC" ), dataset=dataset, model_run_id=train_job.model_run_id,)Below are the first five rows of the predictions table (SMOKERS_DB.PUBLIC.PREDICTIONS_EXPERIMENT_1) on the test data:
| Id | smoking | PROBS | PREDICTED_LABELS |
|---|---|---|---|
| 30249 | 0 | 0.000045 | 0 |
| 18436 | 1 | 0.999944 | 1 |
| 27816 | 0 | 0.000050 | 0 |
| 15941 | 0 | 0.000132 | 0 |
| 7535 | 0 | 0.000040 | 0 |
As expected, the model achieves near-perfect results due to the simplicity of the dataset and the clear distinction between smokers and non-smokers.